ペアエンドシーケンスとシングルリードシーケンスの比較
ペアエンドシーケンスとシングルリードシーケンスの違いは何ですか?
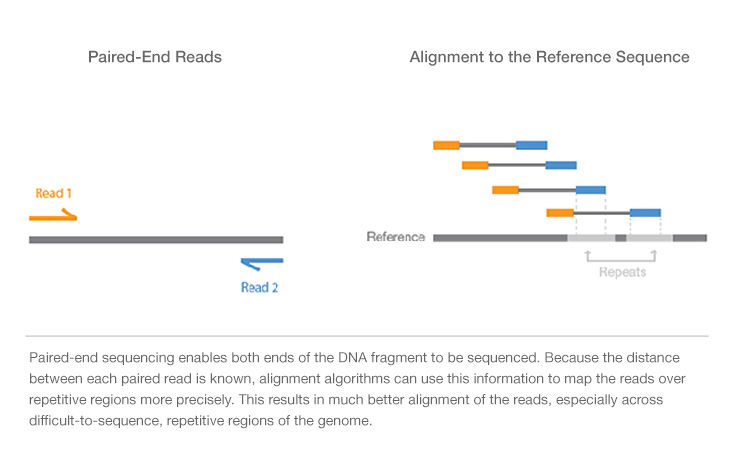
シングルリードシーケンスでは、一端のみからDNAをシーケンスします。これはイルミナのシーケンスを利用する最も簡単な方法です。シングルリードシーケンスとは異なり、ペアエンドシーケンスでは、断片の両端をシーケンスし、高品質でアライメント可能なシーケンスデータを生成できます。ペアエンドシーケンスは、ゲノム再編成や反復配列要素だけでなく、遺伝子融合や新規転写産物の検出を容易にします。
同じ時間と労力のライブラリー調製で2倍のリード数を生成するだけでなく、リードペアとしてアライメントされたシーケンスにより、より正確なリードアライメントが可能になり、シングルリードデータではより困難な挿入欠失(Indel)バリアントを検出することができます1。イルミナの次世代シーケンス(NGS)システムはすべてペアエンドシーケンスが可能です。
ペアエンドシーケンスとシングルエンドシーケンスの利点の比較
ペアエンドシーケンス
- シンプルなペアエンドライブラリー:シンプルなワークフローにより、さまざまなインサートサイズ範囲を生成できます
- 効率的なサンプル使用:シングルリードゲノムDNAまたはcDNAシーケンスと同量のDNAが必要です
- 幅広いアプリケーション:DNAのメチル化や制限消化が不要で、バイサルファイトシーケンシングに使用可能です
- シンプルなデータ解析: ショートインサートライブラリーで高品質のシーケンスアセンブリを可能にします。標準的なシングルリードライブラリー調製プロセスに簡単な変更を加えることで、ペアエンドリード中に各クラスターの順鎖テンプレートと逆鎖テンプレートの両方の鎖を読み取ることを容易にします。どちらのリードにもロングレンジの位置情報が含まれており、リードの高精度なアライメントが可能です。
シングルリードシーケンス
- コスト効率の高い用途:このソリューションは、高品質データを迅速かつ経済的に大量に提供します
- 特定のアプリケーション:シングルリードシーケンスは、Small RNA-Seqやクロマチン免疫沈降シーケンス(ChIP-Seq)などの特定の手法に適しています
ゲノムとトランスクリプトミクスのペアエンドシーケンス
ペアエンドDNAシーケンス
ペアエンドDNAシーケンスリードは、反復シーケンスを含むDNA領域全体で高品質のアライメントを提供し、コンセンサス配列のギャップを埋めることでde novoシーケンス用の長いコンティグを生成します。ペアエンドDNAシーケンスは、挿入、欠失、反転などの一般的なDNA再構成も検出します。
ペアエンドRNAシーケンス
ペアエンドRNAシーケンス(RNA-Seq)は、がんにおける遺伝子融合の検出や、新規スプライスアイソフォームの特性解析などの発見アプリケーションを可能にします。2
ペアエンドRNA-Seqの場合、イルミナは代替の断片化プロトコールを用いたキットを提供し、その後に標準的なイルミナのペアエンドクラスター形成とシーケンスが続きます。
主な製品
Illumina Stranded mRNA Prep
コーディングトランスクリプトームを解析するシンプルでスケーラブル、費用対効果が高く、1日で完了できる迅速ソリューション。
Illumina Stranded Total RNA Prep with Ribo-Zero Plus
コーディングおよびノンコーディングトランスクリプトーム研究用の広範な種類のサンプルから迅速にライブラリーを調製して、非常に優れた柔軟性を持って研究を行うことができます。

遺伝子発現と制御研究による影響力の高い発見
RNAシーケンス技術と、シングルセルRNA-Seqや空間RNA-Seq、タンパク質、クロマチン、メチル化解析などの補完的技術が、生物学と疾患の理解においていかに影響を与えているかをご覧ください。
その他のアプリケーションと手法
イルミナのシーケンスのご紹介
この概要ではシーケンステクノロジーにおける主な進展、主な手法、イルミナシーケンスケミストリーの基本などをご説明します。
NGSシーケンスによるChIPアッセイ
ChIP-Seqを使用して、偏りのないゲノムワイドな遺伝子制御の洞察を得る方法をご覧ください。
DNAシーケンスの方法
さまざまな方法を使用して、DNAシーケンスを小さなターゲット領域、またはゲノム全体に応用する方法をご覧ください。
RNA-Seqの概要
生物学の理解を深めるために、RNA-Seqがいかにトランスクリプトームのコーディング領域およびノンコーディング領域を高解像度で可視化できるかをご覧ください。
シーケンスリード長
適切なシーケンスリード長の選択は、サンプルタイプ、アプリケーション、カバレッジ要件によって異なります。シーケンスランに適したリード長の計算方法を学びましょう。
シングルリードシーケンス
シングルリードシーケンスは、一端のみのDNAシーケンスであり、イルミナシーケンスを利用する最もシンプルな方法です。このソリューションは、迅速かつ経済的に大量の高品質データを提供します。シングルリードシーケンスは、Small RNA-Seqやクロマチン免疫沈降シーケンス(ChIP-Seq)などの特定の手法に適しています。
シーケンシングの実施方法についてさらに知るには:
補足資料
イルミナの合成によるシーケンシング(SBS)
動作中のSBSテクノロジーを見る
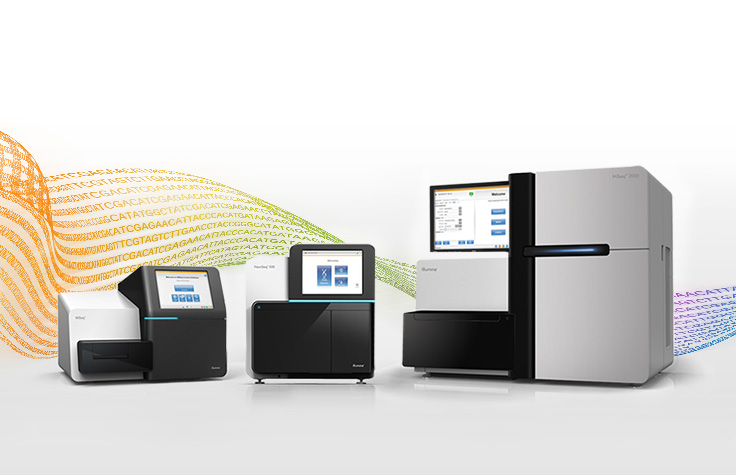

イルミナシーケンスプラットフォーム
ベンチトップおよび生産規模のシーケンサーを確認し、適切なプラットフォームを選択するのに役立つリソースをご利用ください。

ショートペアリードとロングシングルリードの比較
この論文から、同じシーケンス長における長いシングルリードと比較した場合のショートペアリードの利点をご覧ください。

ライブラリー調製
革新的かつ包括的なライブラリー調製ソリューションは、イルミナのシーケンスワークフローの重要な部分です。
参考文献
- Nakazato T, Ohta T, Bono H. Experimental design-based functional mining and characterization of high-throughput sequencing data in the sequence read archive. PLoS One. 2013;8(10):e77910.
- Wang Z, Gerstein M, Snyder M. RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet. 2009;10:57–63.